Faculty & Staff
Dr. Lawrence Stern
 |
assistant PROFESSOR OF CHEMICAL, BIOLOGICAL, AND MATERIALS ENGINEERINGPhone: 813-974-5587 Email: sternl@usf.edu Office: ENG 245 Research cluster: Biotechnology |
|
Honors & Awards
- NSF CAREER Award, 2024
- Ralph E. Powe Junior Faculty Enhancement Award, Oak Ridge Associated Universities, 2021
- USF New Researcher Grant, 2020
- Postdoctoral Fellowship, Tobacco-Related Disease Research Program of the University of California, 2018
Education & Training
- Postdoctoral Fellowship: Beckman Research Institute of the City of Hope
- PhD, Chemical Engineering, University of Minnesota, Twin Cities
- BS, Chemical Engineering, Chemistry, Virginia Tech
Research Overview: Engineering Protein- and Cell-Based Therapies for Disease
The Stern research group aims to understand and engineer better protein- and cell-based therapies by combining high-throughput experimental approaches and computational protein engineering to gain unique insight into sequence-function and structure-function relationships. Major research areas include
- Developing better high-throughput platforms for protein and cell engineering
- Re-engineering natural proteins for therapeutic benefit
- Understanding the building blocks of potent synthetic receptors
Research Description
Synthetic receptors and cell engineering have invigorated the immunotherapy field.
The chimeric antigen receptor (CAR), a synthetic receptor combining the specificity
of an antibody with the cytotoxic capacity of a T cell receptor, has laid the groundwork
for successful engineered cell therapies. While efficacious in a subset of diseases,
limitations of the current CAR platform motivate further study. The Stern Lab aims
to gain a fundamental understanding of the biophysical and biochemical parameters
that govern the efficacy of synthetic receptors while generating clinically relevant
engineering protein- and cell-based therapies.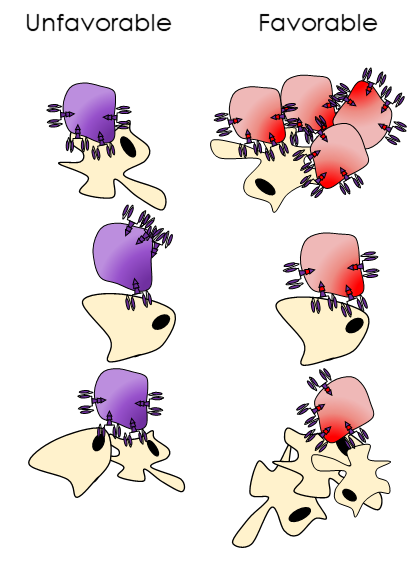
Engineering a better understanding of synthetic receptors: Synthetic receptors, such as CARs or inverted cytokine receptors, are often engineered empirically, fusing natural and engineered domains with minimal optimization. In many cases, this can yield receptors with unfavorable biophysical properties, which can affect receptor activity, expression, and aggregation potential. We aim to combine computational protein design and high-throughput screening with appropriate selective pressures to understand the properties that contribute to receptor fitness and evolve a next generation of improved synthetic receptors.
Redirecting natural molecules for therapeutic benefit: Protein-based inhibitors are a well-established class of drug capable of altering disease states by impeding relevant biology. Often, these drugs are synthetic in nature, requiring several rounds of high-throughput engineering and low-throughput screening to isolate lead candidates. One alternative to this approach is to look to nature, which has generated a wealth of binding molecules with exquisite specificity but activity that is sub-therapeutic. We are applying the principles of structural biology, computational protein engineering approaches, and high-throughput screening to re-engineer natural proteins to act as monotherapies or companions to cell-based therapies.
Developing new high-throughput screening systems for protein engineering: High-throughput screening is a powerful tool for developing new molecules with a desired function, enabling the evaluation of millions to billions of candidate molecules in a single vessel. Successful high-throughput screening is driven by rigorous selective pressure. To date, the most common selective pressure is binding function. We aim to expand high-throughput screening to involve other biologically relevant selective pressures never before explored to adapt this powerful technique to synthetic receptor engineering.