The girls of OCG were able to spend two half days at the Dishaw Lab in USF SP to help identify & explore characteristics of the microbes from a marine organism. The model organism is a simple marine invertebrate — Ciona intestinalis (commonly referred to as a ‘sea squirt’) — which is known to associate with a diverse set of defined microbes referred to as its ‘microbiome.’ Our lab is interested in how symbiotic microbes interact among each other and with the host. In our short time, the girls learned and acquired some hands-on experiences with basic but important methods utilized in microbial ecology and molecular biology.
During our time, the girls performed simple experiments that helped describe ecological (how they behave, how they live) and molecular biological (studies concerning their DNA) principles that define an unknown bacterium isolated from our sea squirt model. The girls were also taught about viruses that infect bacteria, called bacteriophage (or ‘phage’), and how they are able to infect a specific bacterial strain and lyse them. We can visualize this while it’s happening because the bacterial culture, which is very opaque, will start to ‘clear’ over time.
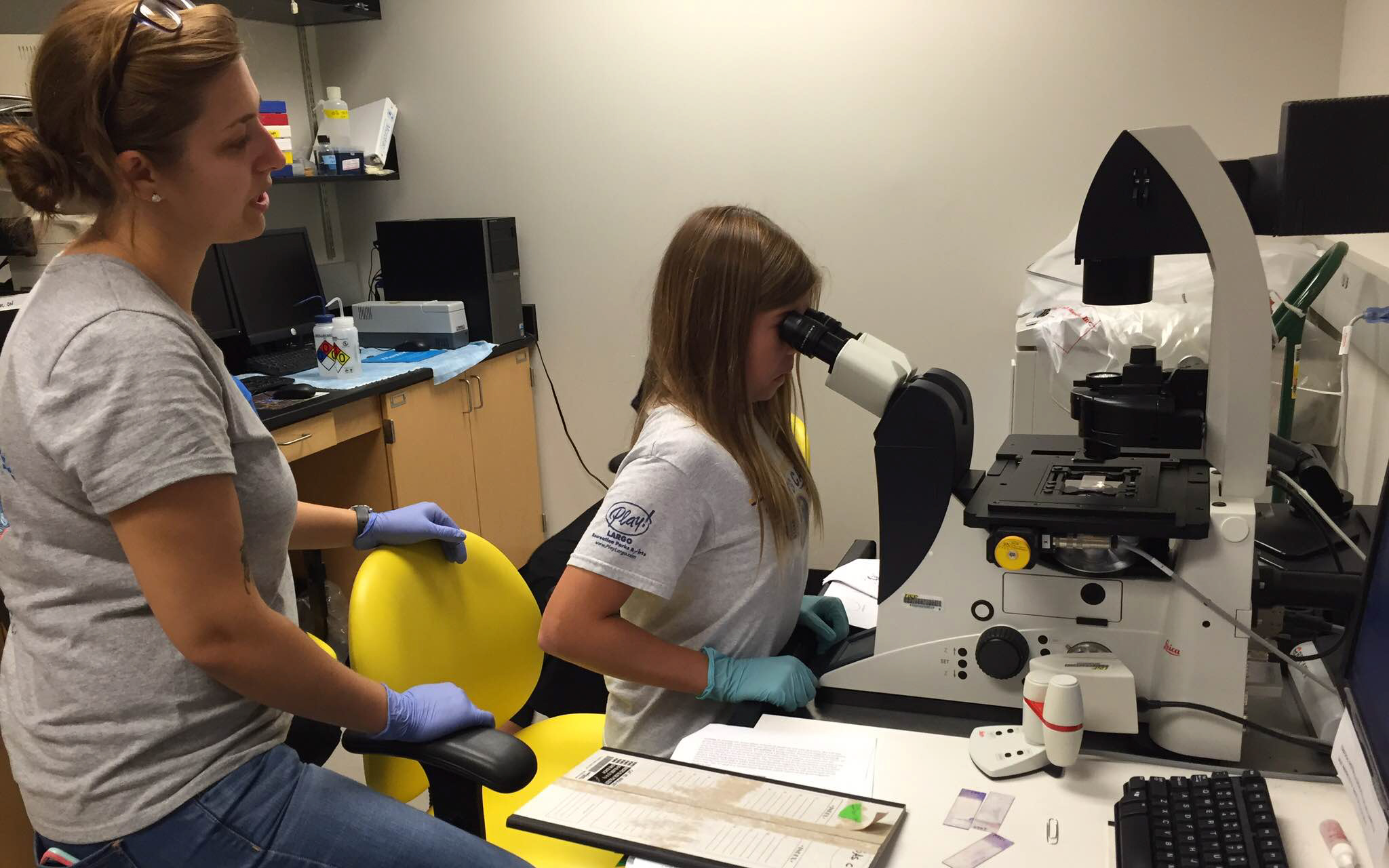
Alexandria Creasy, lab co-leader
We also performed a ‘Gram stain,’ which helps us identify whether our “unknown” is a Gram positive or negative bacterium. This distinction reflects properties of their cell membrane. For example, Gram positive bacteria often have a more rigid capsule that can withstand harsher environmental conditions. This capsule retains the stain and gives the bacteria a much darker bluish/purple color. Our lab also performs experiments where we routinely “colonize” sea squirts with specific bacteria and when we do so, we often use special dyes that allow us to “see” the bacteria as they are taken into the gut. We dyed some bacteria with a fluorescent green dye and watched as the bacteria were taken into the gut of animals and could “see” the bacteria fill the digestive track.
Finally, the girls learned about bacterial fingerprinting. To do this, we used polymerase chain reaction (PCR) to amplify a gene from the bacteria. The amplified gene copies were then cut with special enzymes that recognize a specific DNA sequence. After doing this, the DNA is run on a special gel that separates the “fragments” by size. Because each bacterial species will have a different number of cut sites and in different locations, this generates a unique “fingerprint.” This helps us ultimately identify and keep track of specific bacteria. We think the girls were able to learn some new and very important concepts in our short workshop. And they had fun, just like we did!